Quick Start#
Imports and Paths#
import os, sys
import numpy as np
script_path = os.getcwd()
project_path = os.path.join(script_path, '..')
sys.path.append(project_path)
Installation#
# pip install db-robust-clust
To see the available versions of the package go to the release history at PyPi: https://pypi.org/project/db_robust_clust/#history
Data#
Variable |
Description |
Variable Type |
Possible Categories / Range |
|---|---|---|---|
latitude |
Latitude of the house |
Quantitative |
24.86 - 25.27 |
longitude |
Longitude of the house |
Quantitative |
55.06 - 55.44 |
price |
Market price of the house |
Quantitative |
220000 - 35000000 |
price_per_sqft |
Price per square foot |
Quantitative |
361.87 - 4805.87 |
size in sqft |
Size in square feet |
Quantitative |
294 - 9576 |
no of bedrooms |
Number of bedrooms in the house |
Multiclass |
0, 1, 2, 3, 4, 5 |
no of bathrooms |
Number of bathrooms in the house |
Multiclass |
0, 1, 2, 3, 4, 5, 6 |
quality |
Quality level of the house (response variable) |
Binary |
Low (0), Medium-High-UltraHigh (1) |
balcony |
Indicates if the house has a balcony |
Binary |
true (1), false (0) |
barbecue area |
Indicates if the house has a barbecue area |
Binary |
true (1), false (0) |
private pool |
Indicates if the house has a private pool |
Binary |
true (1), false (0) |
private garden |
Indicates if the house has a private garden |
Binary |
true (1), false (0) |
Note:
The quality variable is the response (target) variable. The remaining variables are mixed-type predictors.
import polars as pl
data_url = "https://raw.githubusercontent.com/FabioScielzoOrtiz/db_robust_clust-docu/refs/heads/main/data/dubai_houses_processed.csv"
df = pl.read_csv(data_url)
response = 'quality'
quant_predictors = ['latitude', 'longitude', 'price', 'size_in_sqft', 'price_per_sqft']
binary_predictors = ['balcony', 'barbecue_area', 'private_pool', 'private_garden']
multiclass_predictors = ['no_of_bedrooms', 'no_of_bathrooms']
y = df[response]
X = df[quant_predictors + binary_predictors + multiclass_predictors]
y.head()
| quality |
|---|
| i64 |
| 1 |
| 1 |
| 1 |
| 0 |
| 1 |
| 1 |
| 1 |
| 1 |
| 0 |
| 1 |
X.head()
| latitude | longitude | price | size_in_sqft | price_per_sqft | balcony | barbecue_area | private_pool | private_garden | no_of_bedrooms | no_of_bathrooms |
|---|---|---|---|---|---|---|---|---|---|---|
| f64 | f64 | i64 | i64 | f64 | i64 | i64 | i64 | i64 | i64 | i64 |
| 25.113208 | 55.138932 | 2700000 | 1079 | 2502.32 | 1 | 1 | 0 | 0 | 1 | 2 |
| 25.106809 | 55.151201 | 2850000 | 1582 | 1801.52 | 1 | 0 | 0 | 0 | 2 | 2 |
| 25.063302 | 55.137728 | 1150000 | 1951 | 589.44 | 1 | 0 | 0 | 0 | 3 | 5 |
| 25.227295 | 55.341761 | 2850000 | 2020 | 1410.89 | 1 | 0 | 0 | 0 | 2 | 3 |
| 25.114275 | 55.139764 | 1729200 | 507 | 3410.65 | 0 | 0 | 0 | 0 | 0 | 1 |
n = len(X)
raw_weights = np.random.rand(n)
simulated_weights = raw_weights / np.sum(raw_weights)
db_robust_clust.models#
p1 = len(quant_predictors)
p2 = len(binary_predictors)
p3 = len(multiclass_predictors)
n_clusters = len(y.unique())
SampleDistClustering#
from db_robust_clust.models import SampleDistClustering
from kmedoids import KMedoids
KMedoids - pam#
kmedoids_method = 'pam'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
frac_sample_size=0.1, metric='euclidean',
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1858, 47], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='ggower', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 1, 1, 1, 1, 1, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1380, 525], dtype=int64))
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
weights = simulated_weights
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
sample_dist_clust.fit(X=X, weights=weights)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='ggower', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 1, 1, 1, 1, 1, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1385, 520], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-7.09e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.47e+01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='relms', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 1, 1, 1, 1, 1, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1382, 523], dtype=int64))
KMedoids - fasterpam#
kmedoids_method = 'fasterpam'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
n_clusters=2,
random_state=123),
frac_sample_size=0.1, metric='euclidean',
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, n_clusters=2, random_state=123)
Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fasterpam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1858, 47], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='ggower', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, n_clusters=2, random_state=123)
Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fasterpam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 1, 1, 1, 1, 1, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1380, 525], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-7.09e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.47e+01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='relms', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, n_clusters=2, random_state=123)
Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fasterpam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 1, 1, 1, 1, 1, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1382, 523], dtype=int64))
KMedoids - alternate#
kmedoids_method = 'alternate'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='alternate',
n_clusters=2,
random_state=123),
frac_sample_size=0.1, metric='euclidean',
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='alternate', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'alternate' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1858, 47], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='alternate',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='ggower', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='alternate', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'alternate' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 1, 1, 1, 1, 1, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1380, 525], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-7.09e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.47e+01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='alternate',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='relms', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='alternate', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'alternate' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 1, 1, 1, 1, 1, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1382, 523], dtype=int64))
KMedoids - fastermsc#
kmedoids_method = 'fastermsc'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='fastermsc',
n_clusters=2,
random_state=123),
frac_sample_size=0.1, metric='euclidean',
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='fastermsc', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fastermsc' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1890, 15], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='fastermsc',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='ggower', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='fastermsc', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fastermsc' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1897, 8], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-7.09e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.47e+01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='fastermsc',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='relms', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='fastermsc', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fastermsc' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1897, 8], dtype=int64))
KMedoids - pamsil#
kmedoids_method = 'pamsil'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pamsil', n_clusters=2,
random_state=123),
frac_sample_size=0.1, metric='euclidean',
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pamsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pamsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 1, 1], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([ 6, 1899], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pamsil', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='ggower', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pamsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pamsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 1, 1], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([ 10, 1895], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-7.09e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.47e+01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pamsil', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='relms', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pamsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pamsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 1, 1], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([ 10, 1895], dtype=int64))
KMedoids - pammedsil#
kmedoids_method = 'pammedsil'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pammedsil',
n_clusters=2,
random_state=123),
frac_sample_size=0.1, metric='euclidean',
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pammedsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pammedsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1890, 15], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
sample_dist_clust.fit(X)
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pammedsil',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='ggower', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pammedsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pammedsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1897, 8], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-7.09e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.47e+01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
SampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pammedsil',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
frac_sample_size=0.1, metric='relms', p1=5, p2=4, p3=2,
random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| frac_sample_size | 0.1 | |
| random_state | 123 | |
| stratify | False | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pammedsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pammedsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0], dtype=uint64)
X_new = X[:10,:]
sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(sample_dist_clust.labels_, return_counts=True)
(array([0, 1], dtype=uint64), array([1897, 8], dtype=int64))
FoldSampleDistClustering#
from db_robust_clust.models import FoldSampleDistClustering
from kmedoids import KMedoids
KMedoids - pam#
kmedoids_method = 'pam'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
metric='euclidean', random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1886, 19], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
meta_frac_sample_size=0.8,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='ggower', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 1, 1])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[1, 0, 0, 1, 0, 0, 0, 0, 0, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([ 554, 1351], dtype=int64))
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
weights = simulated_weights
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
fold_sample_dist_clust.fit(X=X, weights=weights)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='ggower', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 1, 1])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[1, 1, 0, 1, 0, 0, 0, 0, 0, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([ 861, 1044], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
fold_sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.00e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.86e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-1.83e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-3.10e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.03e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.28e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='relms', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 1, 1, 1, 1, 1, 1, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1310, 595], dtype=int64))
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
weights = simulated_weights
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
fold_sample_dist_clust.fit(X=X, weights=weights)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.00e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.86e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-1.83e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-3.10e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.03e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-6.74e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='relms', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 1, 0, 1, 0]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1509, 396], dtype=int64))
KMedoids - fasterpam#
kmedoids_method = 'fasterpam'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
n_clusters=2,
random_state=123),
metric='euclidean', random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, n_clusters=2, random_state=123)
Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fasterpam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1886, 19], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
meta_frac_sample_size=0.8,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='ggower', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, n_clusters=2, random_state=123)
Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fasterpam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 1, 1])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[1, 0, 0, 1, 0, 0, 0, 0, 0, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([ 554, 1351], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
fold_sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.00e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.86e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-1.83e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-3.10e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.03e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.28e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='relms', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, n_clusters=2, random_state=123)
Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fasterpam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 1, 1, 1, 1, 1, 1, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1310, 595], dtype=int64))
KMedoids - alternate#
kmedoids_method = 'alternate'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='alternate',
n_clusters=2,
random_state=123),
metric='euclidean', random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='alternate', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'alternate' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1886, 19], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
meta_frac_sample_size=0.8,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='alternate',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='ggower', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='alternate', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'alternate' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 1, 1])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[1, 0, 0, 1, 0, 0, 0, 0, 0, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([ 554, 1351], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
fold_sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.00e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.86e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-1.83e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-3.10e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.03e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.28e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='alternate',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='relms', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='alternate', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'alternate' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 1, 1, 1, 1, 1, 1, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1310, 595], dtype=int64))
KMedoids - fastermsc#
kmedoids_method = 'fastermsc'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='fastermsc',
n_clusters=2,
random_state=123),
metric='euclidean', random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='fastermsc', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fastermsc' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 1, 1])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([ 19, 1886], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
meta_frac_sample_size=0.8,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='fastermsc',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='ggower', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='fastermsc', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fastermsc' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 1, 0, 0, 0]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1877, 28], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
fold_sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.00e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.86e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-1.83e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-3.10e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.03e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.08e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='fastermsc',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='relms', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='fastermsc', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'fastermsc' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 1, 1, 1, 1, 1, 0]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1596, 309], dtype=int64))
KMedoids - pamsil#
kmedoids_method = 'pamsil'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pamsil',
n_clusters=2,
random_state=123),
metric='euclidean', random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pamsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pamsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1886, 19], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
meta_frac_sample_size=0.8,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pamsil',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='ggower', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pamsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pamsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 0, 1])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[1, 1, 1, 1, 0, 0, 0, 0, 0, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([965, 940], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
fold_sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.00e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.86e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-1.83e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-3.10e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.03e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pamsil',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='relms', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pamsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pamsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 0, 1])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[1, 1, 1, 1, 0, 0, 0, 0, 1, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1076, 829], dtype=int64))
KMedoids - pammedsil#
kmedoids_method = 'pammedsil'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
metric = 'euclidean'#
metric = 'euclidean'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pammedsil',
n_clusters=2,
random_state=123),
metric='euclidean', random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'euclidean' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | None | |
| p2 | None | |
| p3 | None | |
| d1 | None | |
| d2 | None | |
| d3 | None | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pammedsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pammedsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([1, 1, 1, ..., 1, 1, 1])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([ 19, 1886], dtype=int64))
metric = 'ggower'#
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
meta_frac_sample_size=0.8,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pammedsil',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='ggower', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pammedsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pammedsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 0, 0, 1, 0, 0, 0]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1877, 28], dtype=int64))
metric = 'RelMS'#
metric = 'relms'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
fold_sample_dist_clust.fit(X)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.00e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.86e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-1.83e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-3.20e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d2 is not PSD (min eig=-3.10e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-2.03e+00). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\robust_mixed_dist\mixed.py:879: UserWarning: Gram matrix for d3 is not PSD (min eig=-1.08e-01). Transformation applied.
warnings.warn(f'Gram matrix for d{i} is not PSD (min eig={eig_min_val:.2e}). Transformation applied.')
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pammedsil',
n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='relms', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'relms' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pammedsil', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pammedsil' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
fold_sample_dist_clust.labels_
array([0, 0, 0, ..., 0, 0, 0])
X_new = X[:10,:]
fold_sample_dist_clust.predict(X_new)
[0, 0, 0, 0, 1, 1, 1, 1, 1, 0]
np.unique(fold_sample_dist_clust.labels_, return_counts=True)
(array([0, 1]), array([1596, 309], dtype=int64))
db_robust_clust.plots#
clustering_MDS_plot_one_method#
from db_robust_clust.plots import clustering_MDS_plot_one_method
from sklearn.manifold import MDS
from robust_mixed_dist.mixed import generalized_gower_dist_matrix
import seaborn as sns
sns.set_style('whitegrid')
kmedoids_method = 'pam'
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
meta_frac_sample_size=0.8,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
fold_sample_dist_clust.fit(X)
FoldSampleDistClustering(clustering_method=KMedoids(init='build', max_iter=100,
method='pam', n_clusters=2,
random_state=123),
d1='robust_mahalanobis', d2='jaccard', d3='hamming',
metric='ggower', p1=5, p2=4, p3=2, random_state=123)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| clustering_method | KMedoids(init...dom_state=123) | |
| metric | 'ggower' | |
| n_splits | 5 | |
| shuffle | True | |
| random_state | 123 | |
| stratify | False | |
| frac_sample_size | 0.1 | |
| meta_frac_sample_size | 0.8 | |
| p1 | 5 | |
| p2 | 4 | |
| p3 | 2 | |
| d1 | 'robust_mahalanobis' | |
| d2 | 'jaccard' | |
| d3 | 'hamming' | |
| q | 1 | |
| robust_method | 'trimmed' | |
| alpha | 0.05 |
KMedoids(init='build', max_iter=100, method='pam', n_clusters=2,
random_state=123)Parameters
| n_clusters | 2 | |
| metric | 'precomputed' | |
| metric_params | None | |
| method | 'pam' | |
| init | 'build' | |
| max_iter | 100 | |
| random_state | 123 |
mds = MDS(n_components=2, dissimilarity='precomputed', random_state=123)
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
np.random.seed(123)
sample_idx = np.random.choice(range(X.shape[0]), 300)
D = generalized_gower_dist_matrix(
X=X[sample_idx,:], p1=p1, p2=p2, p3=p2,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05, epsilon=0.05, n_iters=20
)
X_mds = mds.fit_transform(D)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\sklearn\manifold\_mds.py:744: FutureWarning: The default value of `n_init` will change from 4 to 1 in 1.9. To suppress this warning, provide some value of `n_init`.
warnings.warn(
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\sklearn\manifold\_mds.py:754: FutureWarning: The default value of `init` will change from 'random' to 'classical_mds' in 1.10. To suppress this warning, provide some value of `init`.
warnings.warn(
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\sklearn\manifold\_mds.py:771: FutureWarning: The `dissimilarity` parameter is deprecated and will be removed in 1.10. Use `metric` instead.
warnings.warn(
clustering_MDS_plot_one_method(X_mds=X_mds, y_pred=fold_sample_dist_clust.labels_[sample_idx],
y_true=None, title="MDS visualization of clustering results",
accuracy=None, time=None,
figsize=(8,7), bbox_to_anchor=(1,1),
title_size=13, title_weight='bold',
points_size=45, title_height=1,
save=False, legend_size=9)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\seaborn\_core\data.py:313: UserWarning: Conversion using Arrow PyCapsule Interface failed due to missing PyArrow>=14 dependency, falling back to (deprecated) interchange protocol. We recommend that you install PyArrow>=14.0.0.
return pd.api.interchange.from_dataframe(data)
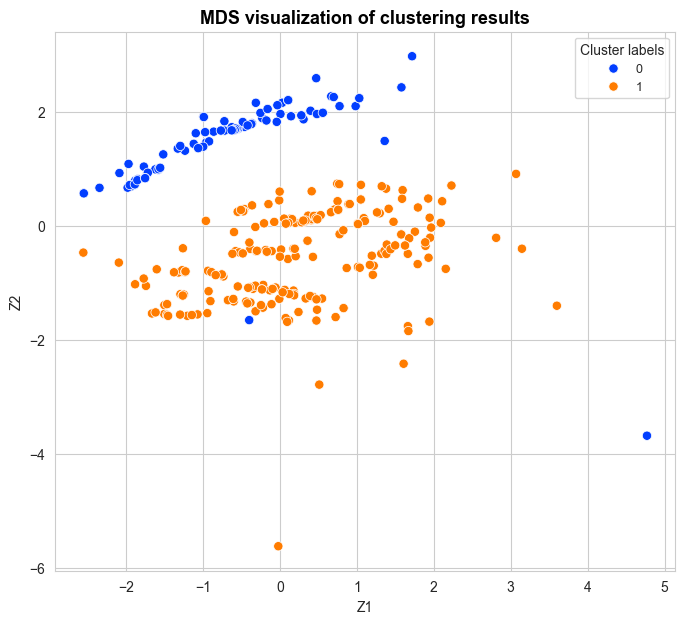
clustering_MDS_plot_multiple_methods#
from db_robust_clust.plots import clustering_MDS_plot_multiple_methods
from db_robust_clust.metrics import adjusted_score
from sklearn.metrics import balanced_accuracy_score
from sklearn.mixture import GaussianMixture
from sklearn.cluster import KMeans
from sklearn_extra.cluster import KMedoids
import time
###############################################################################
kmedoids_method = 'pam'
metric = 'ggower'
d1 = 'robust_mahalanobis'
d2 = 'jaccard'
d3 = 'hamming'
clustering_method = KMedoids(
n_clusters=n_clusters,
metric='precomputed',
method=kmedoids_method,
init='build',
max_iter=100,
random_state=123
)
sample_dist_clust = SampleDistClustering(
clustering_method = clustering_method,
metric = metric,
frac_sample_size=0.1,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05
)
start_time = time.time()
sample_dist_clust.fit(X)
predicted_clusters_sample_dist_clust = sample_dist_clust.labels_
time_fast_kmedoids = time.time() - start_time
###############################################################################
fold_sample_dist_clust = FoldSampleDistClustering(
clustering_method = clustering_method,
metric = metric,
n_splits=5,
shuffle=True,
frac_sample_size=0.1,
meta_frac_sample_size=0.8,
random_state=123,
stratify=False,
p1=p1, p2=p2, p3=p3,
d1=d1, d2=d2, d3=d3,
robust_method='trimmed', alpha=0.05,
)
start_time = time.time()
fold_sample_dist_clust.fit(X=X)
predicted_clusters_fold_sample_dist_clust = fold_sample_dist_clust.labels_
time_fold_fast_kmedoids = time.time() - start_time
###############################################################################
start_time = time.time()
kmeans = KMeans(n_clusters=n_clusters, random_state=123, init='k-means++', n_init='auto', max_iter=300)
kmeans.fit(X)
predicted_clusters_kmeans = kmeans.labels_
time_kmeans = time.time() - start_time
###############################################################################
start_time = time.time()
gmm = GaussianMixture(n_components=n_clusters, random_state=123)
gmm.fit(X)
predicted_clusters_gmm = gmm.predict(X)
time_gmm = time.time() - start_time
###############################################################################
start_time = time.time()
kmedoids = KMedoids(n_clusters=n_clusters, metric='euclidean', method='pam', init='build', max_iter=100, random_state=123)
kmedoids.fit(X)
predicted_clusters_kmedoids = kmedoids.predict(X)
time_kmedoids = time.time() - start_time
###############################################################################
adj_accuracy_sample_dist_clust, adj_predicted_clusters_sample_dist_clust = adjusted_score(y_pred=predicted_clusters_sample_dist_clust, y_true=y, metric=balanced_accuracy_score)
adj_accuracy_fold_sample_dist_clust, adj_predicted_clusters_fold_sample_dist_clust = adjusted_score(y_pred=predicted_clusters_fold_sample_dist_clust, y_true=y, metric=balanced_accuracy_score)
adj_accuracy_kmeans, adj_predicted_clusters_kmeans = adjusted_score(y_pred=predicted_clusters_kmeans, y_true=y, metric=balanced_accuracy_score)
adj_accuracy_gmm, adj_predicted_clusters_gmm = adjusted_score(y_pred=predicted_clusters_gmm, y_true=y, metric=balanced_accuracy_score)
adj_accuracy_kmedoids, adj_predicted_clusters_kmedoids = adjusted_score(y_pred=predicted_clusters_kmedoids, y_true=y, metric=balanced_accuracy_score)
y_pred_dict = {
'SampleDistClust-RobustGGower-KMedoidsPAM': adj_predicted_clusters_sample_dist_clust[sample_idx],
'FoldSampleDistClust-RobustGGower-KMedoidsPAM': adj_predicted_clusters_fold_sample_dist_clust[sample_idx],
'Kmeans': adj_predicted_clusters_kmeans[sample_idx],
'GMM': adj_predicted_clusters_gmm[sample_idx],
'Kmedoids': adj_predicted_clusters_kmedoids[sample_idx]
}
accuracy_dict = {
'SampleDistClust-RobustGGower-KMedoidsPAM': adj_accuracy_sample_dist_clust,
'FoldSampleDistClust-RobustGGower-KMedoidsPAM': adj_accuracy_fold_sample_dist_clust,
'Kmeans': adj_accuracy_kmeans,
'GMM': adj_accuracy_gmm,
'Kmedoids': adj_accuracy_kmedoids,
}
time_dict = {
'SampleDistClust-RobustGGower-KMedoidsPAM': time_fast_kmedoids,
'FoldSampleDistClust-RobustGGower-KMedoidsPAM': time_fold_fast_kmedoids,
'Kmeans': time_kmeans,
'GMM': time_gmm,
'Kmedoids': time_kmedoids,
}
clustering_MDS_plot_multiple_methods(X_mds=X_mds, y_pred=y_pred_dict,
y_true=y[sample_idx],
title="MDS visualization of clustering results",
accuracy=accuracy_dict, time=time_dict, n_rows=2,
figsize=(15,10), bbox_to_anchor=(0.68,-1.9),
title_size=13, subtitles_size=10,
title_weight='bold', points_size=45,
title_height=0.98, legend_size=8,
wspace=0.25, hspace=0.45,
legend_title='Cluster Labels',
n_cols_legend=4, save=False)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\seaborn\_core\data.py:313: UserWarning: Conversion using Arrow PyCapsule Interface failed due to missing PyArrow>=14 dependency, falling back to (deprecated) interchange protocol. We recommend that you install PyArrow>=14.0.0.
return pd.api.interchange.from_dataframe(data)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\seaborn\_core\data.py:313: UserWarning: Conversion using Arrow PyCapsule Interface failed due to missing PyArrow>=14 dependency, falling back to (deprecated) interchange protocol. We recommend that you install PyArrow>=14.0.0.
return pd.api.interchange.from_dataframe(data)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\seaborn\_core\data.py:313: UserWarning: Conversion using Arrow PyCapsule Interface failed due to missing PyArrow>=14 dependency, falling back to (deprecated) interchange protocol. We recommend that you install PyArrow>=14.0.0.
return pd.api.interchange.from_dataframe(data)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\seaborn\_core\data.py:313: UserWarning: Conversion using Arrow PyCapsule Interface failed due to missing PyArrow>=14 dependency, falling back to (deprecated) interchange protocol. We recommend that you install PyArrow>=14.0.0.
return pd.api.interchange.from_dataframe(data)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\seaborn\_core\data.py:313: UserWarning: Conversion using Arrow PyCapsule Interface failed due to missing PyArrow>=14 dependency, falling back to (deprecated) interchange protocol. We recommend that you install PyArrow>=14.0.0.
return pd.api.interchange.from_dataframe(data)
c:\Users\fscielzo\Documents\Proyectos\db_robust_clust-package\.venv\Lib\site-packages\seaborn\_core\data.py:313: UserWarning: Conversion using Arrow PyCapsule Interface failed due to missing PyArrow>=14 dependency, falling back to (deprecated) interchange protocol. We recommend that you install PyArrow>=14.0.0.
return pd.api.interchange.from_dataframe(data)